Tutorial 1: Gradient Descent and AutoGrad#
Week 1, Day 2: Linear Deep Learning
By Neuromatch Academy
Content creators: Saeed Salehi, Vladimir Haltakov, Andrew Saxe
Content reviewers: Polina Turishcheva, Antoine De Comite, Kelson Shilling-Scrivo
Content editors: Anoop Kulkarni, Spiros Chavlis
Production editors: Khalid Almubarak, Gagana B, Spiros Chavlis
Tutorial Objectives#
Day 2 Tutorial 1 will continue on buiding PyTorch skillset and motivate its core functionality: Autograd. In this notebook, we will cover the key concepts and ideas of:
Gradient descent
PyTorch Autograd
PyTorch
nnmodule
Setup#
This a GPU-Free tutorial!
Install and import feedback gadget#
Show code cell source
# @title Install and import feedback gadget
!pip3 install vibecheck datatops --quiet
from vibecheck import DatatopsContentReviewContainer
def content_review(notebook_section: str):
return DatatopsContentReviewContainer(
"", # No text prompt
notebook_section,
{
"url": "https://pmyvdlilci.execute-api.us-east-1.amazonaws.com/klab",
"name": "neuromatch_dl",
"user_key": "f379rz8y",
},
).render()
feedback_prefix = "W1D2_T1"
# Imports
import torch
import numpy as np
from torch import nn
from math import pi
import matplotlib.pyplot as plt
Figure settings#
Show code cell source
# @title Figure settings
import logging
logging.getLogger('matplotlib.font_manager').disabled = True
import ipywidgets as widgets # Interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/content-creation/main/nma.mplstyle")
Plotting functions#
Show code cell source
# @title Plotting functions
from mpl_toolkits.axes_grid1 import make_axes_locatable
def ex3_plot(model, x, y, ep, lss):
"""
Plot training loss
Args:
model: nn.module
Model implementing regression
x: np.ndarray
Training Data
y: np.ndarray
Targets
ep: int
Number of epochs
lss: function
Loss function
Returns:
Nothing
"""
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
ax1.set_title("Regression")
ax1.plot(x, model(x).detach().numpy(), color='r', label='prediction')
ax1.scatter(x, y, c='c', label='targets')
ax1.set_xlabel('x')
ax1.set_ylabel('y')
ax1.legend()
ax2.set_title("Training loss")
ax2.plot(np.linspace(1, epochs, epochs), losses, color='y')
ax2.set_xlabel("Epoch")
ax2.set_ylabel("MSE")
plt.show()
def ex1_plot(fun_z, fun_dz):
"""
Plots the function and gradient vectors
Args:
fun_z: f.__name__
Function implementing sine function
fun_dz: f.__name__
Function implementing sine function as gradient vector
Returns:
Nothing
"""
x, y = np.arange(-3, 3.01, 0.02), np.arange(-3, 3.01, 0.02)
xx, yy = np.meshgrid(x, y, sparse=True)
zz = fun_z(xx, yy)
xg, yg = np.arange(-2.5, 2.6, 0.5), np.arange(-2.5, 2.6, 0.5)
xxg, yyg = np.meshgrid(xg, yg, sparse=True)
zxg, zyg = fun_dz(xxg, yyg)
plt.figure(figsize=(8, 7))
plt.title("Gradient vectors point towards steepest ascent")
contplt = plt.contourf(x, y, zz, levels=20)
plt.quiver(xxg, yyg, zxg, zyg, scale=50, color='r', )
plt.xlabel('$x$')
plt.ylabel('$y$')
ax = plt.gca()
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
cbar = plt.colorbar(contplt, cax=cax)
cbar.set_label('$z = h(x, y)$')
plt.show()
Set random seed#
Executing set_seed(seed=seed) you are setting the seed
Show code cell source
# @title Set random seed
# @markdown Executing `set_seed(seed=seed)` you are setting the seed
# For DL its critical to set the random seed so that students can have a
# baseline to compare their results to expected results.
# Read more here: https://pytorch.org/docs/stable/notes/randomness.html
# Call `set_seed` function in the exercises to ensure reproducibility.
import random
import torch
def set_seed(seed=None, seed_torch=True):
"""
Function that controls randomness. NumPy and random modules must be imported.
Args:
seed : Integer
A non-negative integer that defines the random state. Default is `None`.
seed_torch : Boolean
If `True` sets the random seed for pytorch tensors, so pytorch module
must be imported. Default is `True`.
Returns:
Nothing.
"""
if seed is None:
seed = np.random.choice(2 ** 32)
random.seed(seed)
np.random.seed(seed)
if seed_torch:
torch.manual_seed(seed)
torch.cuda.manual_seed_all(seed)
torch.cuda.manual_seed(seed)
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
print(f'Random seed {seed} has been set.')
# In case that `DataLoader` is used
def seed_worker(worker_id):
"""
DataLoader will reseed workers following randomness in
multi-process data loading algorithm.
Args:
worker_id: integer
ID of subprocess to seed. 0 means that
the data will be loaded in the main process
Refer: https://pytorch.org/docs/stable/data.html#data-loading-randomness for more details
Returns:
Nothing
"""
worker_seed = torch.initial_seed() % 2**32
np.random.seed(worker_seed)
random.seed(worker_seed)
Set device (GPU or CPU). Execute set_device()#
Show code cell source
# @title Set device (GPU or CPU). Execute `set_device()`
# especially if torch modules used.
# inform the user if the notebook uses GPU or CPU.
def set_device():
"""
Set the device. CUDA if available, CPU otherwise
Args:
None
Returns:
Nothing
"""
device = "cuda" if torch.cuda.is_available() else "cpu"
if device != "cuda":
print("GPU is not enabled in this notebook. \n"
"If you want to enable it, in the menu under `Runtime` -> \n"
"`Hardware accelerator.` and select `GPU` from the dropdown menu")
else:
print("GPU is enabled in this notebook. \n"
"If you want to disable it, in the menu under `Runtime` -> \n"
"`Hardware accelerator.` and select `None` from the dropdown menu")
return device
SEED = 2021
set_seed(seed=SEED)
DEVICE = set_device()
Random seed 2021 has been set.
GPU is not enabled in this notebook.
If you want to enable it, in the menu under `Runtime` ->
`Hardware accelerator.` and select `GPU` from the dropdown menu
Section 0: Introduction#
Today, we will go through 3 tutorials.
Starting with Gradient Descent, the workhorse of deep learning algorithms.
The second tutorial will help us build a better intuition about neural networks and basic hyper-parameters.
Finally, in tutorial 3, we learn about the learning dynamics, what the (a good) deep network is learning, and why sometimes they may perform poorly.
Video 0: Introduction#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Introduction_Video")
Section 1: Gradient Descent Algorithm#
Time estimate: ~30-45 mins
Since the goal of most learning algorithms is minimizing the risk (also known as the cost or loss) function, optimization is often the core of most machine learning techniques! The gradient descent algorithm, along with its variations such as stochastic gradient descent, is one of the most powerful and popular optimization methods used for deep learning. Today we will introduce the basics, but you will learn much more about Optimization in the coming days (Week 1, Day 4).
Section 1.1: Gradients & Steepest Ascent#
Video 1: Gradient Descent#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Gradient_Descent_Video")
Before introducing the gradient descent algorithm, let’s review a very important property of gradients. The gradient of a function always points in the direction of the steepest ascent. The following exercise will help clarify this.
Analytical Exercise 1.1: Gradient vector (Optional)#
Given the following function:
find the gradient vector:
Hint: Use the chain rule!
Chain rule: For a composite function \(F(x) = g(h(x)) \equiv (g \circ h)(x)\):
or differently denoted:
Click here for the solution
We can rewrite the function as a composite function:
Using the chain rule:
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Gradient_Vector_Analytical_Exercise")
Coding Exercise 1.1: Gradient Vector#
Implement (complete) the function which returns the gradient vector for \(z=\sin(x^2 + y^2)\).
def fun_z(x, y):
"""
Implements function sin(x^2 + y^2)
Args:
x: (float, np.ndarray)
Variable x
y: (float, np.ndarray)
Variable y
Returns:
z: (float, np.ndarray)
sin(x^2 + y^2)
"""
z = np.sin(x**2 + y**2)
return z
def fun_dz(x, y):
"""
Implements function sin(x^2 + y^2)
Args:
x: (float, np.ndarray)
Variable x
y: (float, np.ndarray)
Variable y
Returns:
Tuple of gradient vector for sin(x^2 + y^2)
"""
#################################################
## Implement the function which returns gradient vector
## Complete the partial derivatives dz_dx and dz_dy
# Complete the function and remove or comment the line below
raise NotImplementedError("Gradient function `fun_dz`")
#################################################
dz_dx = ...
dz_dy = ...
return (dz_dx, dz_dy)
## Uncomment to run
# ex1_plot(fun_z, fun_dz)
We can see from the plot that for any given \(x_0\) and \(y_0\), the gradient vector \(\left[ \dfrac{\partial z}{\partial x}, \dfrac{\partial z}{\partial y}\right]^{\top}_{(x_0, y_0)}\) points in the direction of \(x\) and \(y\) for which \(z\) increases the most. It is important to note that gradient vectors only see their local values, not the whole landscape! Also, length (size) of each vector, which indicates the steepness of the function, can be very small near local plateaus (i.e. minima or maxima).
Thus, we can simply use the aforementioned formula to find the local minima.
In 1847, Augustin-Louis Cauchy used negative of gradients to develop the Gradient Descent algorithm as an iterative method to minimize a continuous and (ideally) differentiable function of many variables.
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Gradient_Vector_Exercise")
Video 2: Gradient Descent - Discussion#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Gradient_Descent_Discussion_Video")
Section 1.2: Gradient Descent Algorithm#
Let \(f(\mathbf{w}): \mathbb{R}^d \rightarrow \mathbb{R}\) be a differentiable function. Gradient Descent is an iterative algorithm for minimizing the function \(f\), starting with an initial value for variables \(\mathbf{w}\), taking steps of size \(\eta\) (learning rate) in the direction of the negative gradient at the current point to update the variables \(\mathbf{w}\).
where \(\eta > 0\) and \(\nabla f (\mathbf{w})= \left( \frac{\partial f(\mathbf{w})}{\partial w_1}, ..., \frac{\partial f(\mathbf{w})}{\partial w_d} \right)\). Since negative gradients always point locally in the direction of steepest descent, the algorithm makes small steps at each point towards the minimum.
Vanilla Algorithm
Inputs: initial guess \(\mathbf{w}^{(0)}\), step size \(\eta > 0\), number of steps \(T\).
For \(t = 0, 1, 2, \dots , T-1\) do
\(\qquad\) \(\mathbf{w}^{(t+1)} = \mathbf{w}^{(t)} - \eta \nabla f \left( \mathbf{w}^{(t)} \right)\)
end
Return: \(\mathbf{w}^{(t+1)}\)
Hence, all we need is to calculate the gradient of the loss function with respect to the learnable parameters (i.e., weights):
Analytical Exercise 1.2: Gradients#
Given \(f(x, y, z) = \tanh \left( \ln \left[1 + z \frac{2x}{sin(y)} \right] \right)\), how easy is it to derive \(\dfrac{\partial f}{\partial x}\), \(\dfrac{\partial f}{\partial y}\) and \(\dfrac{\partial f}{\partial z}\)?
Hint: You don’t have to actually calculate them!
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Gradients_Analytical_Exercise")
Section 1.3: Computational Graphs and Backprop#
Video 3: Computational Graph#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Computational_Graph_Video")
Exercise 1.2 is an example of how overwhelming the derivation of gradients can get, as the number of variables and nested functions increases. This function is still extraordinarily simple compared to the loss functions of modern neural networks. So how can we (as well as PyTorch and similar frameworks) approach such beasts?
Let’s look at the function again:
We can build a so-called computational graph (shown below) to break the original function into smaller and more approachable expressions.
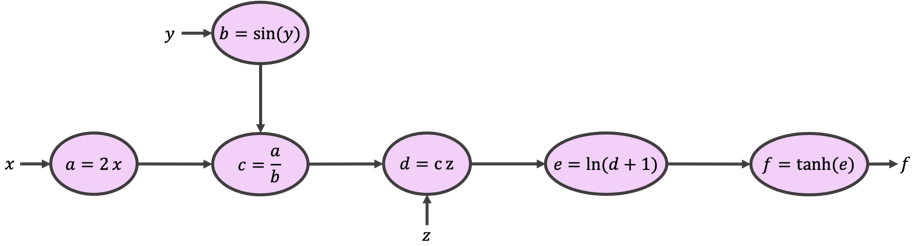
Starting from \(x\), \(y\), and \(z\) and following the arrows and expressions, you would see that our graph returns the same function as \(f\). It does so by calculating intermediate variables \(a,b,c,d,\) and \(e\). This is called the forward pass.
Now, let’s start from \(f\), and work our way against the arrows while calculating the gradient of each expression as we go. This is called the backward pass, from which the backpropagation of errors algorithm gets its name.
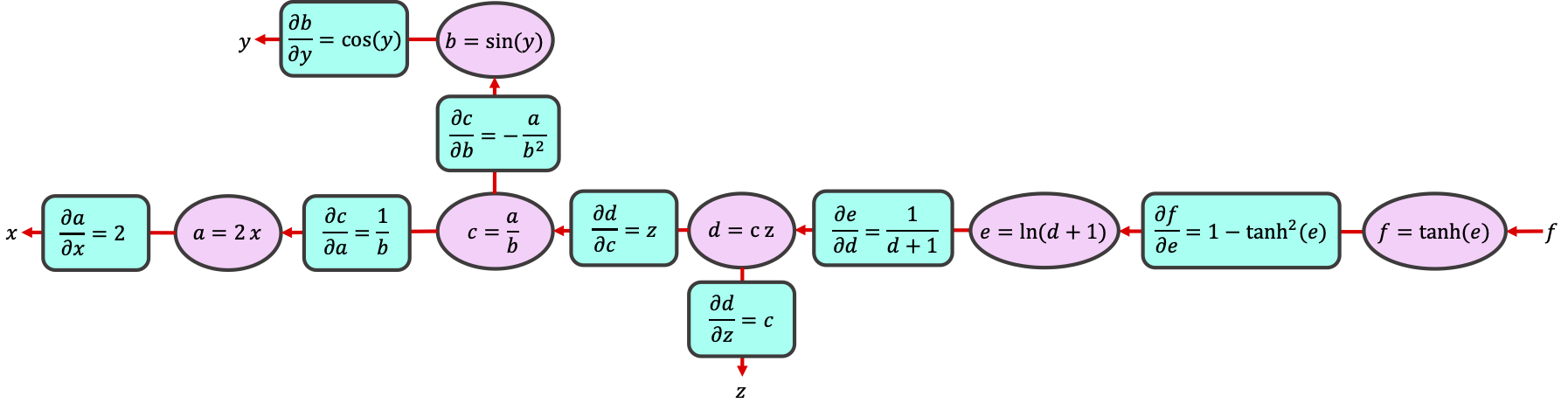
By breaking the computation into simple operations on intermediate variables, we can use the chain rule to calculate any gradient:
Conveniently, the values for \(e\), \(b\), and \(d\) are available to us from when we did the forward pass through the graph. That is, the partial derivatives have simple expressions in terms of the intermediate variables \(a,b,c,d,e\) that we calculated and stored during the forward pass.
Analytical Exercise 1.3: Chain Rule (Optional)#
For the function above, calculate the \(\dfrac{\partial f}{\partial y}\) using the computational graph and chain rule.
Click here for the solution
For more: Calculus on Computational Graphs: Backpropagation
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Chain_Rule_Analytical_Exercise")
Section 2: PyTorch AutoGrad#
Time estimate: ~30-45 mins
Video 4: Auto-Differentiation#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_AutoDifferentiation_Video")
Deep learning frameworks such as PyTorch, JAX, and TensorFlow come with a very efficient and sophisticated set of algorithms, commonly known as Automatic Differentiation. AutoGrad is PyTorch’s automatic differentiation engine. Here we start by covering the essentials of AutoGrad, and you will learn more in the coming days.
Section 2.1: Forward Propagation#
Everything starts with the forward propagation (pass). PyTorch tracks all the instructions, as we declare the variables and operations, and it builds the graph when we call the .backward() pass. PyTorch rebuilds the graph every time we iterate or change it (or simply put, PyTorch uses a dynamic graph).
For gradient descent, it is only required to have the gradients of cost function with respect to the variables we wish to learn. These variables are often called “learnable / trainable parameters” or simply “parameters” in PyTorch. In neural nets, weights and biases are often the learnable parameters.
Coding Exercise 2.1: Buiding a Computational Graph#
In PyTorch, to indicate that a certain tensor contains learnable parameters, we can set the optional argument requires_grad to True. PyTorch will then track every operation using this tensor while configuring the computational graph. For this exercise, use the provided tensors to build the following graph, which implements a single neuron with scalar input and output.
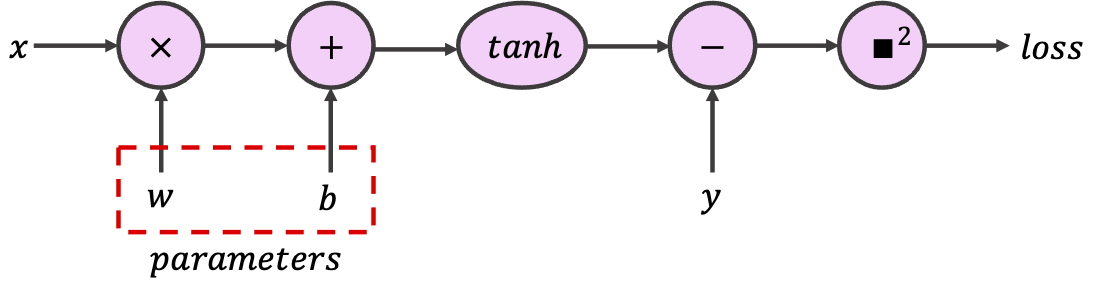
class SimpleGraph:
"""
Implementing Simple Computational Graph
"""
def __init__(self, w, b):
"""
Initializing the SimpleGraph
Args:
w: float
Initial value for weight
b: float
Initial value for bias
Returns:
Nothing
"""
assert isinstance(w, float)
assert isinstance(b, float)
self.w = torch.tensor([w], requires_grad=True)
self.b = torch.tensor([b], requires_grad=True)
def forward(self, x):
"""
Forward pass
Args:
x: torch.Tensor
1D tensor of features
Returns:
prediction: torch.Tensor
Model predictions
"""
assert isinstance(x, torch.Tensor)
#################################################
## Implement the the forward pass to calculate prediction
## Note that prediction is not the loss, but the value after `tanh`
# Complete the function and remove or comment the line below
raise NotImplementedError("Forward Pass `forward`")
#################################################
prediction = ...
return prediction
def sq_loss(y_true, y_prediction):
"""
L2 loss function
Args:
y_true: torch.Tensor
1D tensor of target labels
y_prediction: torch.Tensor
1D tensor of predictions
Returns:
loss: torch.Tensor
L2-loss (squared error)
"""
assert isinstance(y_true, torch.Tensor)
assert isinstance(y_prediction, torch.Tensor)
#################################################
## Implement the L2-loss (squred error) given true label and prediction
# Complete the function and remove or comment the line below
raise NotImplementedError("Loss function `sq_loss`")
#################################################
loss = ...
return loss
feature = torch.tensor([1]) # Input tensor
target = torch.tensor([7]) # Target tensor
## Uncomment to run
# simple_graph = SimpleGraph(-0.5, 0.5)
# print(f"initial weight = {simple_graph.w.item()}, "
# f"\ninitial bias = {simple_graph.b.item()}")
# prediction = simple_graph.forward(feature)
# square_loss = sq_loss(target, prediction)
# print(f"for x={feature.item()} and y={target.item()}, "
# f"prediction={prediction.item()}, and L2 Loss = {square_loss.item()}")
It is important to appreciate the fact that PyTorch can follow our operations as we arbitrarily go through classes and functions.
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Building_a_computational_graph_Exercise")
Section 2.2: Backward Propagation#
Here is where all the magic lies. In PyTorch, Tensor and Function are interconnected and build up an acyclic graph, that encodes a complete history of computation. Each variable has a grad_fn attribute that references a function that has created the Tensor (except for Tensors created by the user - these have None as grad_fn). The example below shows that the tensor c = a + b is created by the Add operation and the gradient function is the object <AddBackward...>. Replace + with other single operations (e.g., c = a * b or c = torch.sin(a)) and examine the results.
a = torch.tensor([1.0], requires_grad=True)
b = torch.tensor([-1.0], requires_grad=True)
c = a + b
print(f'Gradient function = {c.grad_fn}')
Gradient function = <AddBackward0 object at 0x7f29c3589a60>
For more complex functions, printing the grad_fn would only show the last operation, even though the object tracks all the operations up to that point:
print(f'Gradient function for prediction = {prediction.grad_fn}')
print(f'Gradient function for loss = {square_loss.grad_fn}')
Now let’s kick off the backward pass to calculate the gradients by calling .backward() on the tensor we wish to initiate the backpropagation from. Often, .backward() is called on the loss, which is the last node on the graph. Before doing that, let’s calculate the loss gradients by hand:
Where \(y_t\) is the target (true label), and \(y_p\) is the prediction (model output). We can then compare it to PyTorch gradients, which can be obtained by calling .grad on the relevant tensors.
Important Notes:
Learnable parameters (i.e.
requires_gradtensors) are “contagious”. Let’s look at a simple example:Y = W @ X, whereXis the feature tensors andWis the weight tensor (learnable parameters,requires_grad), the newly generated output tensorYwill be alsorequires_grad. So any operation that is applied toYwill be part of the computational graph. Therefore, if we need to plot or store a tensor that isrequires_grad, we must first.detach()it from the graph by calling the.detach()method on that tensor..backward()accumulates gradients in the leaf nodes (i.e., the input nodes to the node of interest). We can call.zero_grad()on the loss or optimizer to zero out all.gradattributes (see autograd.backward for more information).Recall that in python we can access variables and associated methods with
.method_name. You can use the commanddir(my_object)to observe all variables and associated methods to your object, e.g.,dir(simple_graph.w).
# Analytical gradients (Remember detaching)
ana_dloss_dw = - 2 * feature * (target - prediction.detach())*(1 - prediction.detach()**2)
ana_dloss_db = - 2 * (target - prediction.detach())*(1 - prediction.detach()**2)
square_loss.backward() # First we should call the backward to build the graph
autograd_dloss_dw = simple_graph.w.grad # We calculate the derivative w.r.t weights
autograd_dloss_db = simple_graph.b.grad # We calculate the derivative w.r.t bias
print(ana_dloss_dw == autograd_dloss_dw)
print(ana_dloss_db == autograd_dloss_db)
References and more#
Section 3: PyTorch’s Neural Net module (nn.Module)#
Time estimate: ~30 mins
Video 5: PyTorch nn module#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Pytorch_nn_module_Video")
PyTorch provides us with ready-to-use neural network building blocks, such as layers (e.g., linear, recurrent, etc.), different activation and loss functions, and much more, packed in the torch.nn module. If we build a neural network using torch.nn layers, the weights and biases are already in requires_grad mode and will be registered as model parameters.
For training, we need three things:
Model parameters: Model parameters refer to all the learnable parameters of the model, which are accessible by calling
.parameters()on the model. Please note that NOT all therequires_gradtensors are seen as model parameters. To create a custom model parameter, we can usenn.Parameter(A kind of Tensor that is to be considered a module parameter).Loss function: The loss that we are going to be optimizing, which is often combined with regularization terms (coming up in few days).
Optimizer: PyTorch provides us with many optimization methods (different versions of gradient descent). Optimizer holds the current state of the model and by calling the
step()method, will update the parameters based on the computed gradients.
You will learn more details about choosing the right model architecture, loss function, and optimizer later in the course.
Section 3.1: Training loop in PyTorch#
We use a regression problem to study the training loop in PyTorch.
The task is to train a wide nonlinear (using \(\tanh\) activation function) neural net for a simple \(\sin\) regression task. Wide neural networks are thought to be really good at generalization.
Generate the sample dataset#
Show code cell source
# @markdown #### Generate the sample dataset
set_seed(seed=SEED)
n_samples = 32
inputs = torch.linspace(-1.0, 1.0, n_samples).reshape(n_samples, 1)
noise = torch.randn(n_samples, 1) / 4
targets = torch.sin(pi * inputs) + noise
plt.figure(figsize=(8, 5))
plt.scatter(inputs, targets, c='c')
plt.xlabel('x (inputs)')
plt.ylabel('y (targets)')
plt.show()
Random seed 2021 has been set.
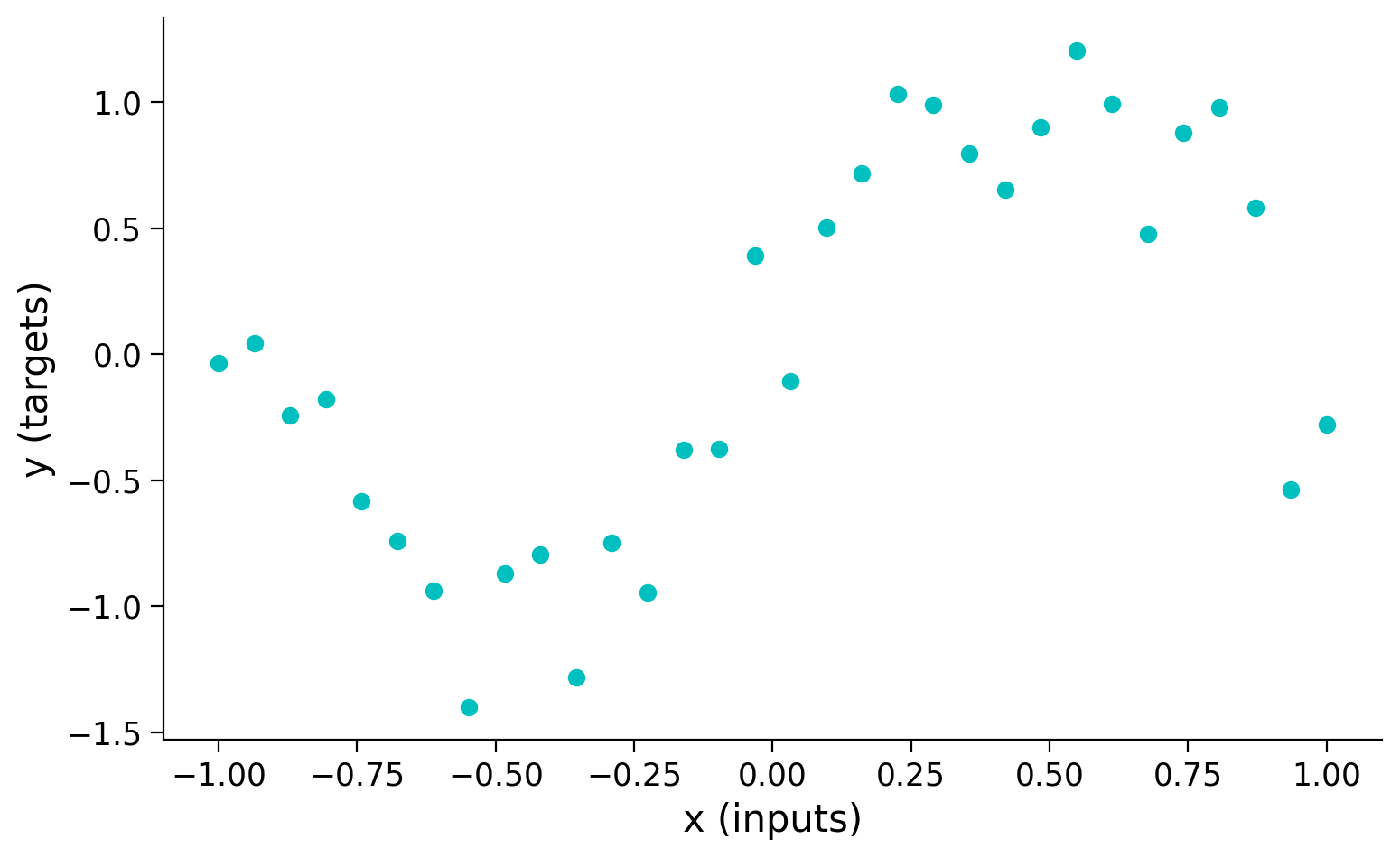
Let’s define a very wide (512 neurons) neural net with one hidden layer and nn.Tanh() activation function.
class WideNet(nn.Module):
"""
A Wide neural network with a single hidden layer
Structure is as follows:
nn.Sequential(
nn.Linear(1, n_cells) + nn.Tanh(), # Fully connected layer with tanh activation
nn.Linear(n_cells, 1) # Final fully connected layer
)
"""
def __init__(self):
"""
Initializing the parameters of WideNet
Args:
None
Returns:
Nothing
"""
n_cells = 512
super().__init__()
self.layers = nn.Sequential(
nn.Linear(1, n_cells),
nn.Tanh(),
nn.Linear(n_cells, 1),
)
def forward(self, x):
"""
Forward pass of WideNet
Args:
x: torch.Tensor
2D tensor of features
Returns:
Torch tensor of model predictions
"""
return self.layers(x)
We can now create an instance of our neural net and print its parameters.
# Creating an instance
set_seed(seed=SEED)
wide_net = WideNet()
print(wide_net)
Random seed 2021 has been set.
WideNet(
(layers): Sequential(
(0): Linear(in_features=1, out_features=512, bias=True)
(1): Tanh()
(2): Linear(in_features=512, out_features=1, bias=True)
)
)
# Create a mse loss function
loss_function = nn.MSELoss()
# Stochstic Gradient Descent optimizer (you will learn about momentum soon)
lr = 0.003 # Learning rate
sgd_optimizer = torch.optim.SGD(wide_net.parameters(), lr=lr, momentum=0.9)
The training process in PyTorch is interactive - you can perform training iterations as you wish and inspect the results after each iteration.
Let’s perform one training iteration. You can run the cell multiple times and see how the parameters are being updated and the loss is reducing. This code block is the core of everything to come: please make sure you go line-by-line through all the commands and discuss their purpose with your pod.
# Reset all gradients to zero
sgd_optimizer.zero_grad()
# Forward pass (Compute the output of the model on the features (inputs))
prediction = wide_net(inputs)
# Compute the loss
loss = loss_function(prediction, targets)
print(f'Loss: {loss.item()}')
# Perform backpropagation to build the graph and compute the gradients
loss.backward()
# Optimizer takes a tiny step in the steepest direction (negative of gradient)
# and "updates" the weights and biases of the network
sgd_optimizer.step()
Loss: 0.888942301273346
Coding Exercise 3.1: Training Loop#
Using everything we’ve learned so far, we ask you to complete the train function below.
def train(features, labels, model, loss_fun, optimizer, n_epochs):
"""
Training function
Args:
features: torch.Tensor
Features (input) with shape torch.Size([n_samples, 1])
labels: torch.Tensor
Labels (targets) with shape torch.Size([n_samples, 1])
model: torch nn.Module
The neural network
loss_fun: function
Loss function
optimizer: function
Optimizer
n_epochs: int
Number of training iterations
Returns:
loss_record: list
Record (evolution) of training losses
"""
loss_record = [] # Keeping recods of loss
for i in range(n_epochs):
#################################################
## Implement the missing parts of the training loop
# Complete the function and remove or comment the line below
raise NotImplementedError("Training loop `train`")
#################################################
... # Set gradients to 0
predictions = ... # Compute model prediction (output)
loss = ... # Compute the loss
... # Compute gradients (backward pass)
... # Update parameters (optimizer takes a step)
loss_record.append(loss.item())
return loss_record
set_seed(seed=2021)
epochs = 1847 # Cauchy, Exercices d'analyse et de physique mathematique (1847)
## Uncomment to run
# losses = train(inputs, targets, wide_net, loss_function, sgd_optimizer, epochs)
# ex3_plot(wide_net, inputs, targets, epochs, losses)
Random seed 2021 has been set.
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Training_loop_Exercise")
Summary#
In this tutorial, we covered one of the most basic concepts of deep learning; the computational graph and how a network learns via gradient descent and the backpropagation algorithm. We have seen all of these using PyTorch modules and we compared the analytical solutions with the ones provided directly by the PyTorch module.
Video 6: Tutorial 1 Wrap-up#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Tutorial_1_WrapUp_Video")

